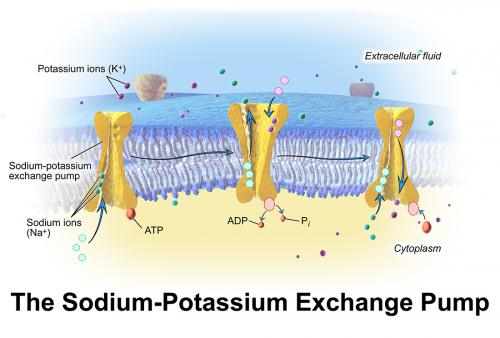
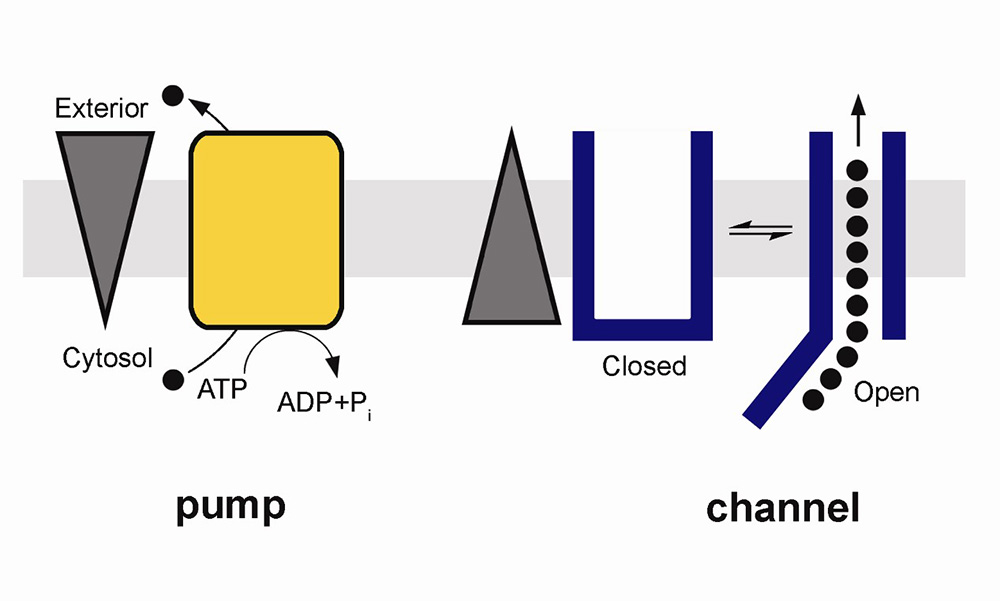
Pumps, channels and transporters are huge molecules. The Na+,K+-ATPase, or sodium pump, the first ion-transporting enzyme to be discovered, consists of three subunits with a total molecular weight of around 150 000 g mol–1. For most chemists, the idea of tackling such a molecule and developing an understanding of how it functions or how it could be manipulated (e.g. inhibited) would seem very daunting. Nevertheless, chemists have much to contribute to research in this field and one can expect that their important contributions will increase as time passes. But first, what are pumps, channels and transporters and what do they do?
Pumps, channels and transporters are all proteins embedded in the membranes of living cells or cell organelles. Their role is to provide a mechanism for the transport of ions or small molecules across the membrane.
Kidney function is just one example where pumps, channels and transporters play a crucial role. Another is in nerve and muscle function (including the heart). The electrical impulses that travel along nerve cells to muscle cells and eventually stimulate muscle contraction are triggered by the opening and closing of voltage-gated ion channels in the plasma membrane of these cells. The signalling agent for muscle contraction and relaxation is in fact Ca2+, whose concentration in the cytoplasm of muscle cells is controlled by a complex interplay of Ca2+-dependent channels, pumps and transporters. Even such a simple thing as cell volume is controlled by ion transport. Without the continual pumping of Na+ and K+ ions across the plasma membrane of all animal cells by the sodium pump, the osmotic conditions across the membrane could change and a cell would be in danger of swelling or even bursting.
Any research field goes through different stages, with the skills of different types of scientists being required at different times as new knowledge accumulates and the field matures. In the case of membrane transport, as is the case of many fields, the first stage was discovery. The first isolation of an ion-transporting membrane protein, the sodium pump, was achieved by Jens Christian Skou of the University of Aarhus, Denmark, who in 1957 purified the protein from crab nerve. From research carried out by physiologists predominantly in the US in the late 1930s and 1940s, it had already become clear that animal cells must possess a sodium pump to counteract the passive Na+ permeability of the plasma membrane. For his discovery, Skou received the Nobel Prize in Chemistry in 1997. Why did it take the Nobel Committee 40 years to recognise his work? One reason was probably scepticism, particularly within the chemical community, that a protein could distinguish between two ions, Na+ and K+, that have such similar physical and chemical properties. Indeed, the mechanism by which proteins accomplish this is still unresolved and is the subject of much current research. Another reason for the late recognition of Skou’s discovery was probably that much further research was required before there could be no more doubt that the protein he’d isolated and purified actually incorporated the complete active transport machinery necessary for sodium and potassium ions.
For the 30 years following Skou’s discovery, research in the membrane transport field was dominated by medical scientists, biochemists and physiologists, in particular electrophysiologists. In the early 1960s, Robert Crane suggested that energy could be stored in living cells via ion concentration gradients, which could be utilised for nutrient transport, as described in the case of the Na+, glucose transporter (see box, p. 22). This idea was taken further by Peter Mitchell, who developed his chemiosmotic theory describing how an H+ gradient could be coupled to ATP synthesis in mitochondria, for which he received the 1978 Nobel Prize in Chemistry.
This exciting initial discovery phase of the field continued, with the first ion channels and transporters being isolated in the 1970s. Throughout this period, however, from the late 1950s to the early 1980s, nothing was known about the molecular structure of pumps, channels and transporters, apart from the fact that they were proteins, i.e. polypeptides. Nevertheless, once these molecules had been isolated and purified, much could be learned about their substrate specificity and their reaction mechanisms by both steady-state and pre-steady kinetic methods, e.g. using radioactive isotope flux measurements, current recordings via electrophysiological techniques and rapid reaction techniques such as stopped-flow with spectroscopic detection. Such techniques led to the development of mechanisms, with letters and numbers signifying different protein conformations. For example, for the P-type family of ion pumps, to which the sodium pump belongs, the mechanism is known as the Albers–Post or E1–E2 cycle, with the E1 and E2 conformations alternating between phosphorylated (E1P and E2P) and unphosphorylated states.
In the 1980s, the membrane transport field started to gradually become more molecular. Following major advances in the field of genetic engineering and molecular biology, the amino acid sequences of pumps, channels and transporters could begin to be determined. The first sequences of the sodium pump and the sarcoplasmic reticulum Ca2+-ATPase were published in Nature in 1985. Today, countless sequences of membrane proteins are stored in databases such as that of the National Center for Biotechnology Information (US Library of Medicine, NIH) and are freely available as a resource to scientists all around the world.
A further significant advance was the development of methods of crystallisation of membrane proteins in detergent-solubilised form and the determination of their three-dimensional structures at atomic resolution by X-ray crystallography. The first structure to be determined was that of a bacterial photosynthetic reaction centre, for which Michel, Deisenhofer and Huber won the Nobel Prize in Chemistry in 1988. Ten years after the publication of this first structure, one could still count the number of published crystal structures of membrane proteins on two hands, but, although crystallisation is still not easy, structures are now appearing at increasingly faster rates. In 2003, Roderick MacKinnon won the Nobel Prize in Chemistry for determining the first crystal structure of an ion channel, and crystal structures of a number of ion pumps (including the sodium pump and the sarcoplasmic reticulum Ca2+-ATPase) in different conformational states have now appeared. The E1–E2 cycle is still used in describing the mechanism of these pumps, but now actual molecular structures can be associated with the letters and numbers.
With the expansion of the structural information available, computational chemists and physicists have more recently been entering the field. Simultaneously with developments in molecular and structural biology, the capability of molecular dynamics simulations to handle larger systems, i.e. more atoms, and to extend simulations over longer timescales has been continually improving. Although still not simple, with appropriate caution and careful consideration of the systems, it is now possible to tackle simulations of membrane proteins embedded in a simulated membrane and to consider questions such as the origin of ion specificity or the mechanism of ion permeation through a channel.
Apart from playing crucial physiological roles, pumps, channels and transporters are also intimately involved in disease and are potential targets in the treatment of disease. Diseases associated with defects in ion channels are often referred to as ‘channelopathies’. Because of the involvement of ion channels in nerve excitability, there are a number of ion channel defects that cause epilepsy. As far as I am aware, no one has suggested the word ‘pumpopathies’ for pump-associated diseases, but probably simply because it is impossible to pronounce. In fact, many diseases associated with defects in ion pumps do exist. An example is Wilson’s disease, which is associated with an increase in the concentration of copper ions within the cytoplasm due to a defect in a Cu+-ATPase responsible for pumping copper ions out of the cell.
When discussing disease, it is important to realise that pumps, channels and transporters are not exclusively animal proteins. They are present in all forms of life. Bacteria and plants possess an H+-ATPase in their membranes, and utilise the energy stored in the H+ gradient for the absorption of nutrients, in a similar way to the utilisation of the Na+ gradient by animal cells. Inhibition of crucial ion pumps of infectious microbial or bacterial organisms is, thus, potentially a powerful strategy for combatting disease. An important example is malaria, which is caused by the protozoan Plasmodium falciparum. It has recently been discovered that after infection of a host’s red blood cells, the Plasmodium expresses a sodium pump, PfATP4, in its plasma membrane in order to counterbalance Na+ influx from the cytoplasm of the dying host cell. Inhibition of PfATP4, thus, represents a potentially novel antimalarial treatment. In 2013, the group of Professor Kiaran Kirk at ANU reported that the new antimalarial drug cipargamin, a spiroindolone drug, acts by inhibition of PfATP4. With the growing international problem of antimicrobial resistance to current antibiotics, such new strategies are urgently required. In the case of malaria, resistance to current treatments involving the use of the drug artemisinin have already been reported throughout South-East Asia.
Many infectious bacteria have recently been found to express a calcium pump in their membranes after infecting the host. Bacterial cells contain much lower calcium concentrations than their animal hosts. Therefore, they require calcium pumps to counterbalance the influx of Ca2+ from the host and prevent the bacterial cell swelling and bursting. This is analogous to the Na+ influx problem encountered by Plasmodium just described. Inhibition of bacterial calcium pumps could, thus, represent an effective treatment for a multitude of bacterial infections, including pneumonia and anthrax.
With the increasing level of structural information on pumps, channels and transporters becoming available in the form of amino acid sequences and crystal structures, and the more sophisticated molecular understanding of their mechanisms that this is yielding, the hope is of more informed predictions of the structures of small molecules that could act as inhibitors. This is an area in which both synthetic and theoretical chemists will increasingly be in demand. Returning to the problem of malaria as a prime example, although cipargamin is an effective inhibitor of PfATP4, like many keys opening a single door, a bewildering range of drugs with varying structures have been found to have inhibitory activity. Because no crystal structure of PfATP4 has yet been determined, it is not currently clear how these promising new drugs work. The Open Source Malaria Consortium founded by my colleague Associate Professor Mat Todd at the University of Sydney is examining one such potential antimalarial and has decided to use crowdsourcing (with a little cash incentive) to try and solve this problem once and for all (bit.ly/2kXZYap). Here is an excellent example of how collaborations between chemists, bioinformaticians, molecular biologists and parasitologists are being used to reveal the role of ion pumps in the treatment of human disease.